Overview
MIP is an AI-native research platform for biology. It combines a domain-specialized reasoning engine with real-time access to 40+ scientific databases, a live literature index, and a containerized compute environment — giving you a single surface to ask questions, analyze data, run pipelines, and generate structured outputs across the full breadth of biological research. The platform is built around three layers: a reasoning layer that understands biology at the level of genes, proteins, pathways, variants, drugs, and disease mechanisms; a data layer that connects to authoritative public databases and your own uploaded datasets; and a compute layer that can execute code, run autonomous multi-step pipelines, and produce publication-quality results.MIP is a research and clinical decision-support tool. All outputs are intended to assist qualified professionals and should be reviewed by a domain expert before being used in any clinical or regulatory context.
Who MIP is for
MIP is built for researchers, clinicians, and teams working across biology and biomedicine. This includes:- Molecular and cell biologists investigating gene function, protein interactions, and disease mechanisms
- Clinical geneticists and genetic counselors interpreting variants and generating reports
- Bioinformaticians running pipelines, validating data, and exploring multi-omics datasets
- Drug discovery scientists mapping targets, pathways, and compound activity
- Translational researchers bridging bench findings with clinical and population data
- Lab directors and principal investigators overseeing analysis across projects and team members
Architecture
MIP’s architecture is designed to keep reasoning, data, and compute tightly integrated so you never have to context-switch between tools.Reasoning layer
A domain-specialized AI engine that understands biological entities and relationships — genes, variants, proteins, pathways, diseases, drugs, phenotypes. It decomposes complex questions into tool calls, searches the right databases, interprets results, and synthesizes answers with citations. Multiple specialized agents (genomics, literature, structural biology, pharmacology) are composed dynamically based on your question.
Data layer
Real-time connections to 40+ public databases (see full list below), your uploaded datasets (VCFs, expression matrices, custom files), and a live PubMed/preprint literature index. The reasoning layer queries these sources on every request — results are never cached or stale.
Compute layer
A containerized execution environment with Python and R, pre-installed scientific libraries (pandas, NumPy, Biopython, scanpy, pydeseq2, matplotlib, and 15+ others), and support for long-running background jobs. Code runs in isolated sandboxes with results streamed back inline or available asynchronously.
Genomic variant analysis
MIP includes a full variant interpretation workflow for clinical and research genomics:| Step | What happens |
|---|---|
| Data ingestion | Upload VCF files in GRCh37 (hg19) or GRCh38 (hg38) format. The platform auto-detects the genome assembly and validates input. |
| Annotation | Variants are annotated using Ensembl VEP 115 with RefSeq transcripts, population frequencies from gnomAD, clinical classifications from ClinVar, and in silico predictions from SIFT, PolyPhen-2, and AlphaMissense. See the Annotation Pipeline for technical details. |
| Classification | Each variant receives an automated ACMG/AMP pathogenicity classification based on 28 standard criteria. Classifications can be refined interactively using AI-enhanced evidence gathering. |
| Interpretation | Annotated variants are available for querying, filtering, and analysis through the AI chat interface, variant browser, and code execution environment. |
| Reporting | Structured clinical reports can be generated on demand with findings, ACMG evidence, and case summaries. |
| Application | Description |
|---|---|
| Whole Exome Sequencing (WES) | Coding regions, typically 30,000-100,000 variants |
| Whole Genome Sequencing (WGS) | Full genome coverage, up to 1 million variants |
| Targeted gene panels | Custom or curated gene sets, typically 100-5,000 variants |
| Chromosomal Microarray Analysis (CMA) | Copy number variant detection from array-based platforms |
Multi-omics and custom data
Beyond genomics, MIP supports analysis of any biological dataset you bring:- Transcriptomics — RNA-seq count matrices, differential expression, pathway enrichment
- Proteomics — Protein quantification, structure exploration via AlphaFold and PDB
- Metabolomics — Metabolite identification, pathway mapping
- Single-cell — scanpy-based workflows for clustering, trajectory analysis, marker identification
- Custom datasets — Upload CSVs, TSVs, or other tabular data and analyze with natural language or code
Data sources
MIP integrates evidence from 40+ public databases across six categories. All sources are queried in real time.| Database | Category | Used for |
|---|---|---|
| PubMed | Literature | Peer-reviewed publications, systematic reviews, case reports |
| bioRxiv / medRxiv | Literature | Preprints and emerging research |
| ClinVar | Clinical | Variant clinical significance, review status, conditions |
| OMIM | Clinical | Gene-disease relationships and inheritance patterns |
| ClinGen | Clinical | Gene-disease validity and dosage sensitivity |
| Orphanet | Clinical | Rare disease classifications and epidemiology |
| gnomAD | Population | Allele frequencies across global populations |
| dbSNP | Population | Variant rsID identifiers |
| GWAS Catalog | Population | Genome-wide association study results |
| Ensembl VEP | Annotation | Consequence prediction, transcript mapping, HGVS nomenclature |
| RefSeq | Annotation | Transcript definitions and gene models |
| UniProt | Protein | Protein sequence, function, domains, PTMs, active sites |
| PDB | Structure | Experimentally determined protein structures |
| AlphaFold | Structure | AI-predicted protein structures |
| AlphaMissense | Prediction | Deep learning-based missense pathogenicity scores |
| SIFT | Prediction | Sequence homology-based functional impact |
| PolyPhen-2 | Prediction | Structure and sequence-based functional impact |
| ChEMBL | Drug discovery | Bioactivity data, compound-target relationships |
| Open Targets | Drug discovery | Target-disease associations, tractability |
| DrugBank | Pharmacology | Drug mechanisms, interactions, pharmacokinetics |
| PharmGKB | Pharmacogenomics | Drug-gene interactions and dosing guidelines |
| KEGG | Pathways | Metabolic and signaling pathway maps |
| Reactome | Pathways | Curated biological pathway knowledge base |
| Gene Ontology | Annotation | Functional annotation, biological process, molecular function |
| MyGene.info | Gene | Gene metadata aggregation across multiple sources |
| NCBI Gene | Gene | Gene summaries, nomenclature, orthologs |
| HPO | Phenotype | Human Phenotype Ontology terms and gene associations |
| Phen2Gene | Phenotype | Phenotype-to-gene matching with ranked scores |
Output types
MIP produces structured, versionable outputs from any analysis:Variant annotations and classifications
Variant annotations and classifications
HGVS nomenclature, consequence terms, transcript mapping, population frequencies, in silico predictions, and automated ACMG/AMP classifications with per-criterion evidence. See ACMG Classification.
Clinical reports
Clinical reports
Variant interpretation reports, findings summaries, case overviews, and ACMG evidence reports. See Clinical Reports.
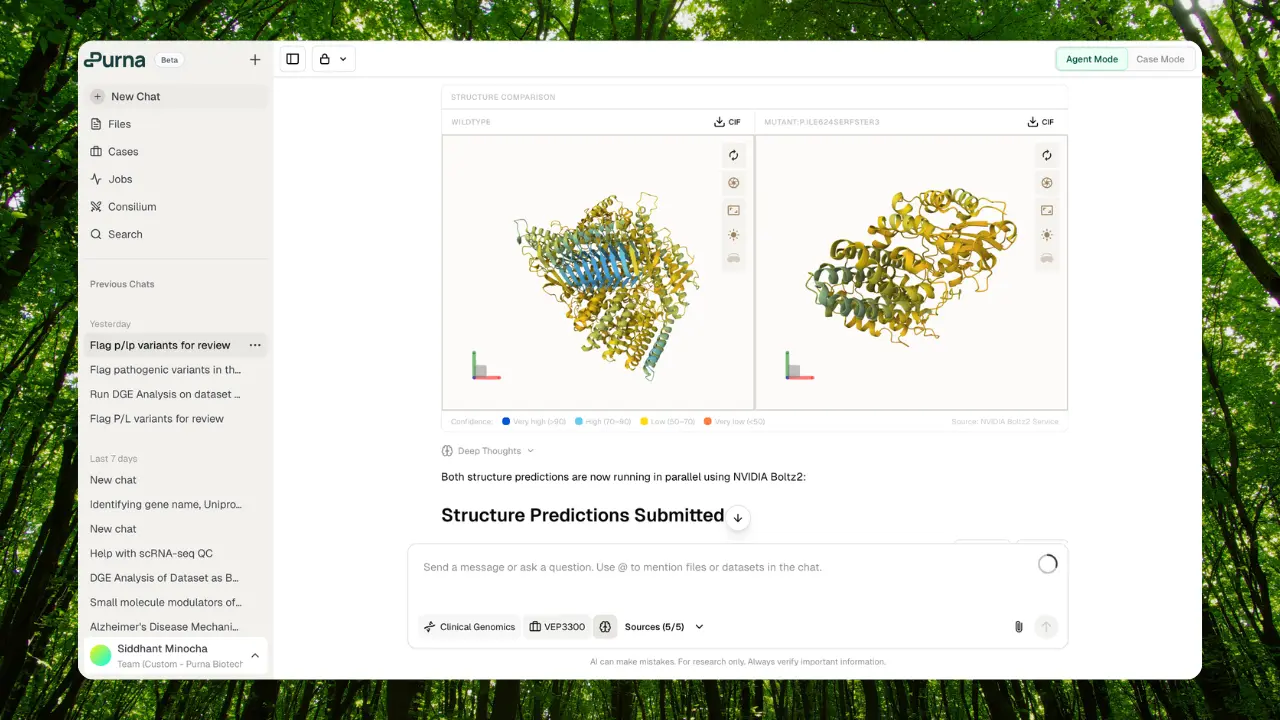
Protein visualizations
Protein visualizations
Color-coded protein sequence viewers with domain annotations, PTMs, active sites, transmembrane regions, and feature tracks. Single protein and comparison modes with sequence alignment.
Literature synthesis
Literature synthesis
Summarized findings from PubMed searches, contradiction analysis, evidence tables, and citation-backed answers to specific research questions.
Hypothesis evaluation
Hypothesis evaluation
Multi-agent debate outputs including refined hypotheses, agent consensus, unresolved tensions, minority dissent, and aggregated literature evidence. See Consilium.
Visualizations and charts
Visualizations and charts
Publication-quality figures generated from natural language or code — bar, line, scatter, pie, heatmaps, volcano plots, and more.
Code execution results
Code execution results
Tables, statistical outputs, and programmatic analysis from Python or R running in a sandboxed compute environment. See Code Execution.
Spreadsheets and exports
Spreadsheets and exports
Tabular data in CSV or spreadsheet format, downloadable and shareable.
Limitations
- MIP is a decision-support tool. It does not make autonomous clinical decisions and is not intended to replace the judgment of qualified professionals.
- Automated ACMG/AMP classifications are rule-based starting points. All classifications should be reviewed by a trained professional before clinical use.
- Variant annotations reflect the versions of reference databases at the time of processing. Re-annotation may be needed as databases are updated.
- AI-generated analysis should be verified against primary sources. MIP provides citations to make this straightforward.
Security and privacy
- All data is encrypted in transit (TLS) and at rest.
- Patient and research data is stored in your organization’s isolated environment and is never shared across accounts.
- Access to individual cases and projects is controlled through role-based permissions configurable in Team settings.
- All data access and mutations are logged to an immutable audit trail.
Never include patient-identifying information such as names, medical record numbers, or contact details in case names, notes, or shared conversations. Use anonymized identifiers for all cases.
