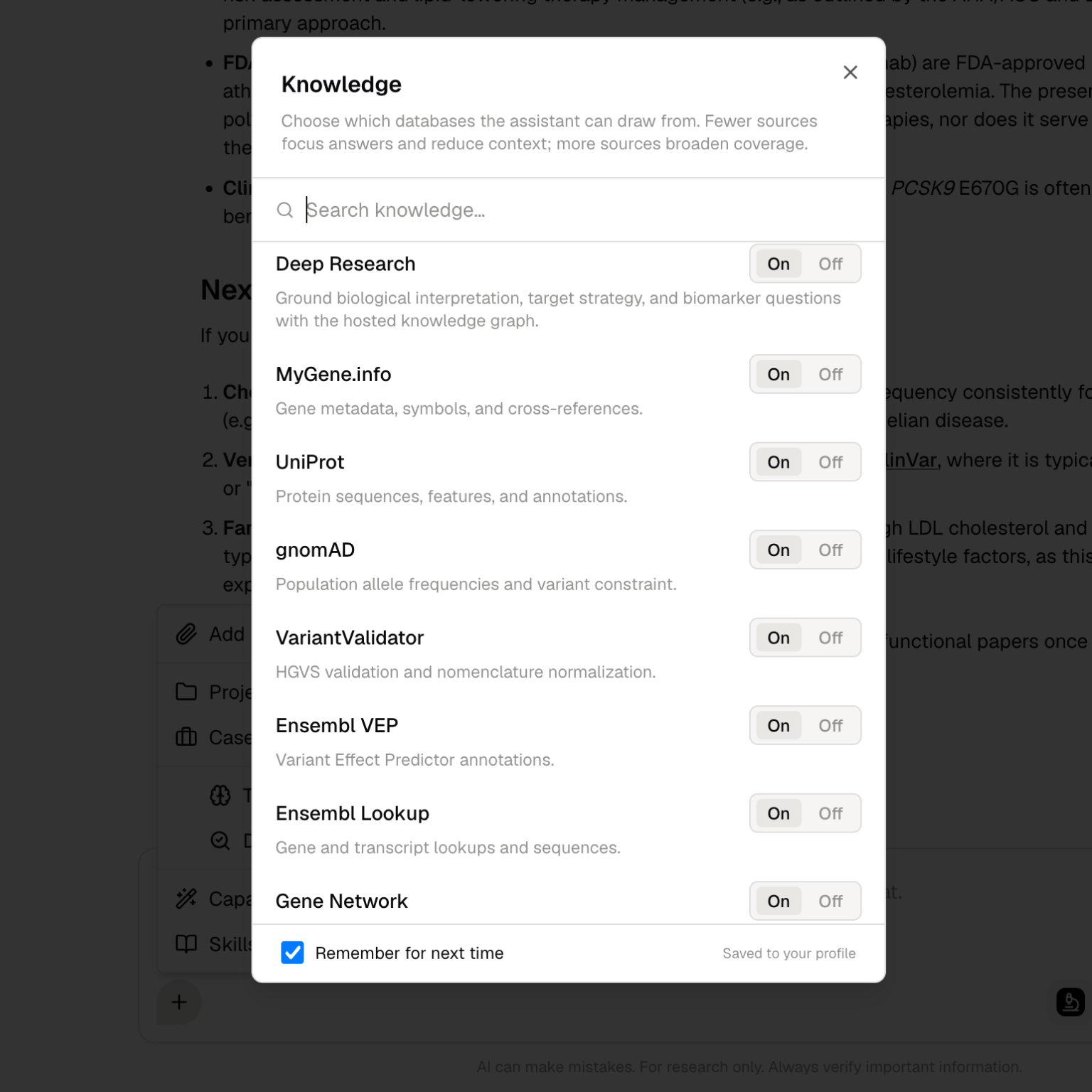
What Deep Research does
Deep Research helps you investigate biological questions across literature, databases, and connected research context. It can help with:- Literature review and source comparison
- Gene, variant, disease, pathway, protein, and compound research
- Mechanism exploration
- Target and hypothesis assessment
- Evidence summaries for documents or team review
- Follow-up questions that build on the same research thread
Start a Deep Research run
- Open a chat in Purna.
- Ask a research question that needs source-backed investigation.
- Select or request Deep Research when the question requires structured retrieval and synthesis.
- Review the plan, sources, and final answer.
- Ask follow-up questions or move the synthesis into Scribe Lab.
- “Run Deep Research on the evidence linking PCSK9 inhibition to familial hypercholesterolemia outcomes.”
- “Compare recent evidence for MAPK pathway reactivation as a resistance mechanism in melanoma.”
- “Investigate whether this target has tractability, pathway, disease, and literature support.”
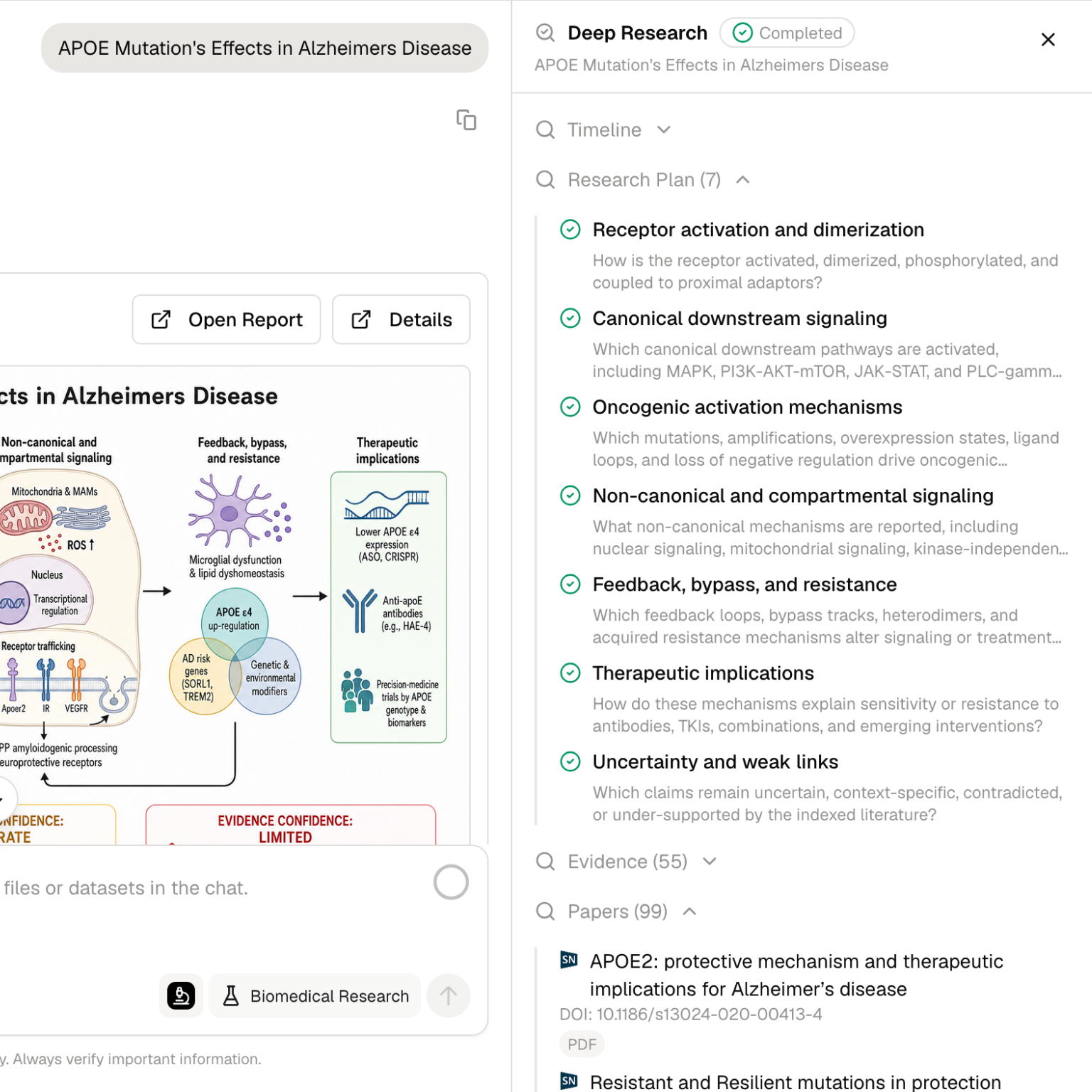
How it works
Deep Research follows a structured workflow:- Plan - Break the question into subtopics, source types, and decision criteria.
- Retrieve - Search relevant literature, databases, and biological context.
- Compare - Evaluate claims across sources and identify agreements, contradictions, and limitations.
- Synthesize - Produce a research answer with cited evidence and clear caveats.
- Continue - Support follow-up questions in the same research context.
Sources and citations
Deep Research can use high-authority biological sources such as literature indexes, pathway databases, gene and variant resources, protein databases, compound resources, and disease knowledge sources. When reviewing an answer:- Check the cited sources attached to key claims.
- Look for caveats and limitations.
- Ask follow-up questions when a source conflict is unclear.
- Move only the relevant conclusions into a document or report.
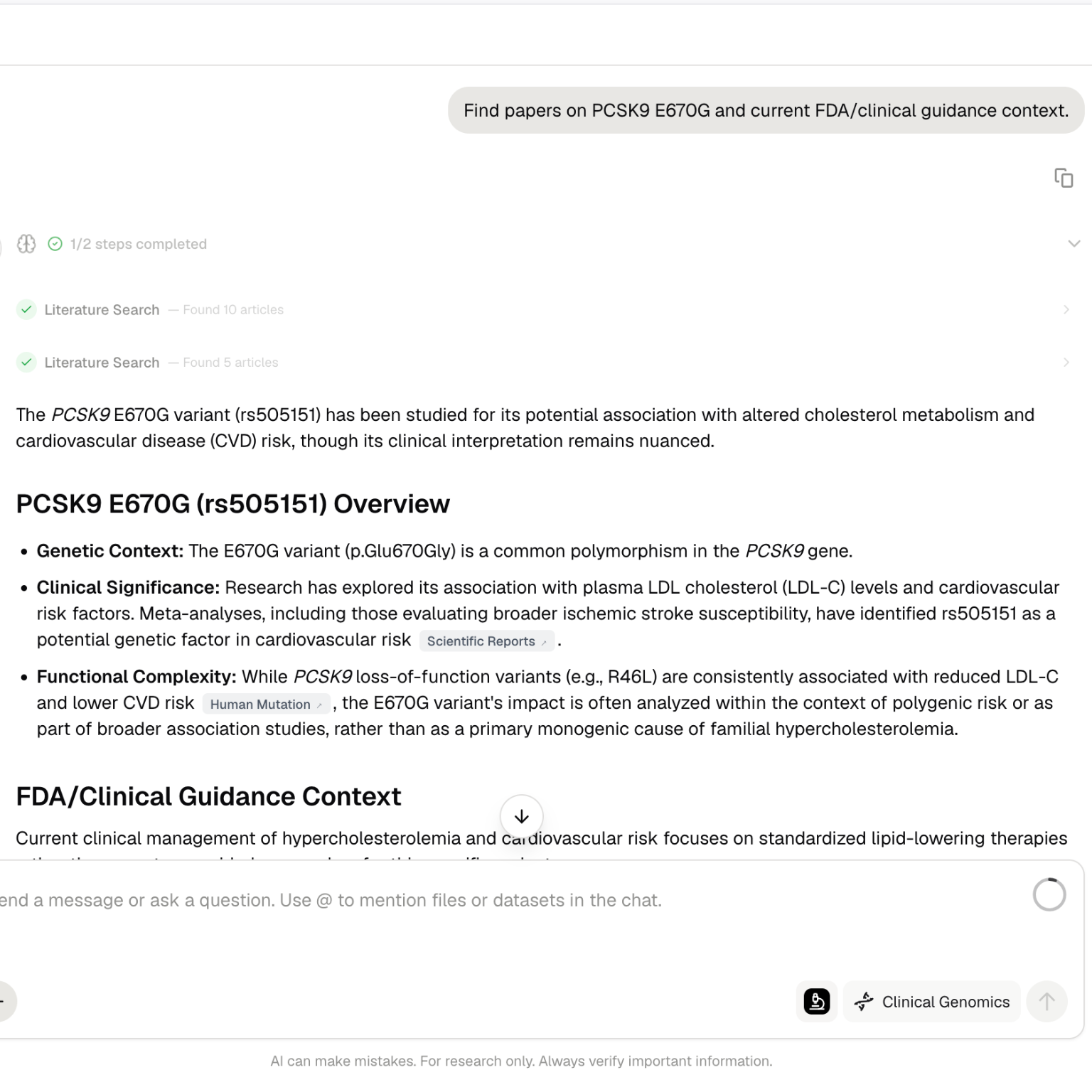
Purna Graph context
Deep Research can use connected biological context from Purna Graph. This helps relate entities such as genes, variants, diseases, proteins, pathways, compounds, and publications. This is useful when the question is not limited to one source. For example, a target assessment may require literature evidence, pathway context, disease associations, structural information, and compound activity.Work with Scribe Lab
Use Scribe Lab when a Deep Research answer needs to become a durable document. A common workflow is:- Run Deep Research on the question.
- Review the sources and synthesis.
- Ask for a structured outline or brief.
- Move the material into Scribe Lab.
- Edit the document and preserve source-backed claims.
Best practices
- Ask a focused research question.
- Include the organism, disease area, gene, pathway, or target when relevant.
- Specify whether you want a table, brief, comparison, or citation-backed narrative.
- Ask Deep Research to call out uncertainty and conflicting evidence.
- Use follow-up questions to narrow broad findings.
Deep Research supports source-grounded investigation, but it does not replace expert scientific review. Verify important claims before making research, clinical, regulatory, or investment decisions.
