MIP can run two families of protein language models on your sequences, directly from chat:
- ESM-2 — for generic proteins. Use it for embeddings or per-position mutation likelihoods on enzymes, receptors, scaffolds, or any non-antibody protein.
- AbLang-2 — for paired antibodies. Use it on heavy + light chain pairs for antibody-aware embeddings or per-position mutation likelihoods that reflect natural antibody repertoires.
Both capabilities return a live, inline result card in chat, store the full numerical output as a downloadable dataset, and bill on actual compute time.
What each capability does
Embeddings
An embedding is a fixed-length vector that represents a full protein sequence. Two sequences with similar embeddings are treated as similar by the underlying model, typically reflecting similar fold, family, or function.
Use embeddings when you want to:
- Cluster a set of sequences by similarity
- Compare two or more proteins numerically
- Search a library for sequences similar to a reference
- Build features for downstream ML (activity prediction, property prediction)
You submit a batch of sequences, and the card returns a compact summary of how many were encoded and the vector dimensionality. The full numerical embeddings are saved as a downloadable JSON dataset so you can load them into a notebook, a clustering pipeline, or your own models.
Mutation likelihoods
A mutation likelihood is a per-position score for every possible amino acid substitution at every residue. High scores at a position mean the model finds multiple alternatives plausible, which is often a signal of an evolvable, tolerant, or flexible site. Low scores mean the model strongly prefers the natural residue.
Use mutation likelihoods when you want to:
- Identify candidate sites for mutagenesis or affinity maturation
- Flag unusual residues in a sequence
- Rank substitutions at a known hotspot
- Get a starting point for protein engineering, before committing to wet-lab rounds
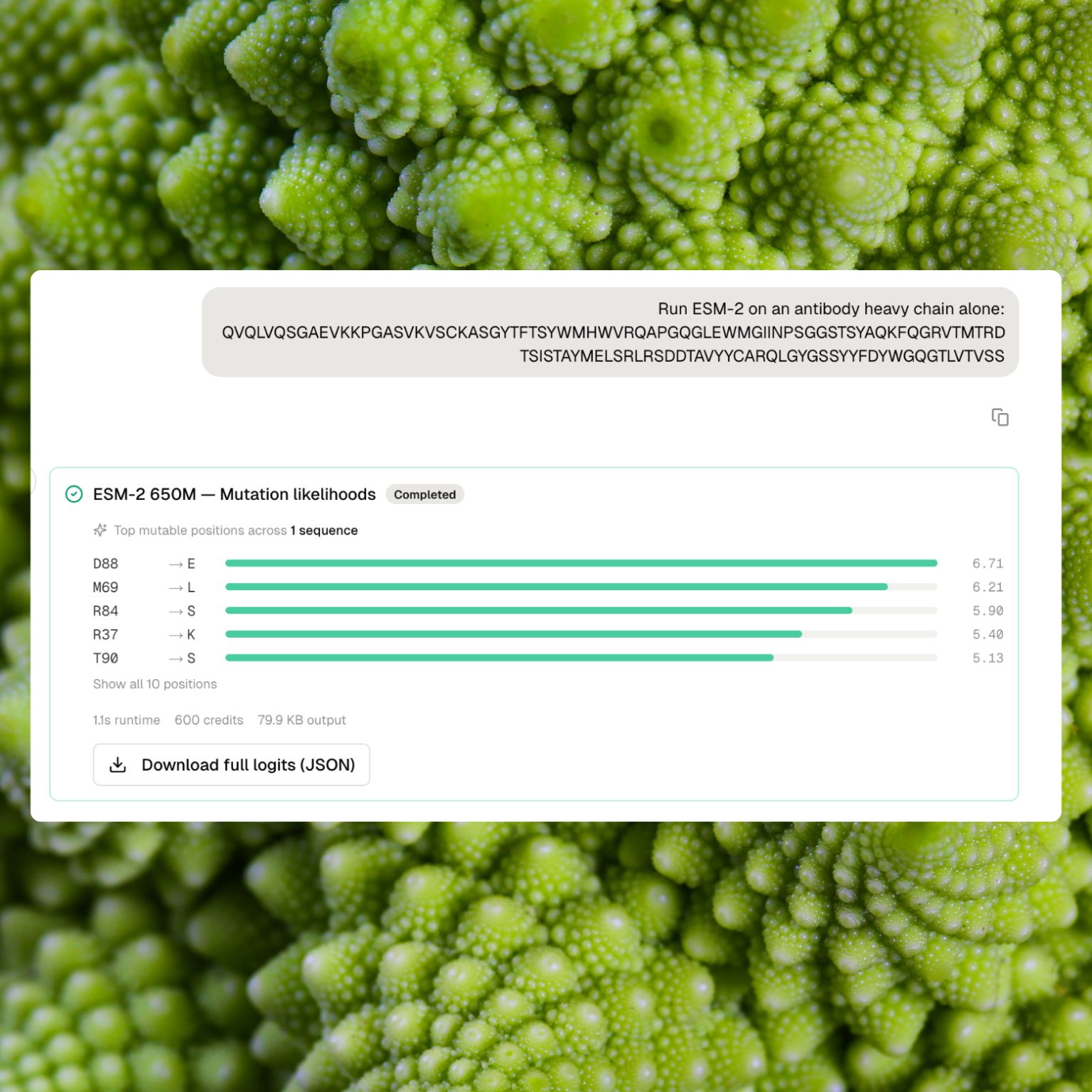
The result card shows the top mutable positions with their highest-likelihood substitutions in a compact list. The full per-position × 20-amino-acid likelihood matrix is saved as a downloadable dataset for deeper analysis.
When to use ESM-2 vs AbLang-2
| You are working with | Use |
|---|
| Enzymes, receptors, scaffold proteins, generic non-antibody proteins | ESM-2 |
| A paired antibody (heavy + light chain) | AbLang-2 |
| Only a heavy chain, and you want an antibody-aware signal | ESM-2 is acceptable; AbLang-2 requires a pair |
| A large protein (up to 2,048 residues) | ESM-2 |
| Multiple antibodies for a comparative embedding run | AbLang-2 |
Asking for a run
You can trigger either capability with a natural-language prompt. A few examples:
ESM-2 embeddings
- “Compute ESM-2 embeddings for this sequence: MKWV…”
- “Encode these 5 protein sequences with ESM-2 so I can cluster them.”
ESM-2 mutation likelihoods
- “Give me per-position mutation likelihoods for this enzyme using ESM-2.”
- “Run ESM-2 mutation scoring on this protein and show me the top mutable sites.”
AbLang-2 embeddings
- “Encode these heavy/light chain pairs with AbLang-2.”
- “Give me AbLang-2 embeddings for this antibody pair.”
AbLang-2 mutation likelihoods
- “Run AbLang-2 mutation likelihoods on this paired antibody.”
- “Show me the top mutable CDR positions for this antibody pair.”
MIP validates the input, submits the job, and shows a live result card in the chat. You do not need to wait or refresh — the card updates itself as the job moves from queued to running to completed.
ESM-2
| Field | Requirement |
|---|
| Items per run | 1 to 8 sequences |
| Length | 1 to 2,048 residues per sequence |
| Alphabet | Extended IUPAC amino acids and - for gaps (ACDEFGHIKLMNPQRSTVWYBJOUXZ) |
AbLang-2
| Field | Requirement |
|---|
| Items per run | 1 to 32 paired antibodies |
| Length | 1 to 1,024 residues per chain |
| Alphabet | Standard 20 amino acids |
| Pairing | Each item must include both heavy and light chains |
The result card
When you submit, an inline card appears in the chat. It updates in place without a page refresh.
While running: a compact status card shows the model, the action, and a spinner. You can continue the chat — the card tracks its own job.
When complete:
- Embeddings runs show the number of sequences encoded and the vector dimensionality (for example,
3 × 1280 dims for ESM-2 or 2 × 480 dims for AbLang-2), plus a one-line hint on common uses.
- Mutation-likelihood runs show a ranked list of the top mutable positions with their highest-likelihood substitutions and scores. Positions are labeled with the observed residue and a clear arrow to the predicted alternative (for example,
Q127 → E (3.02)). The list starts with the top 5 and expands on request.
On failure: the card shows the failure reason (invalid input, provider error, out of credits) and the run is not billed for the failed attempt.
Downloading the full output
Both capabilities save their full numerical output as a downloadable dataset, accessible from:
- The result card’s Download action
- The Files view, under your datasets
Embedding runs save the full vector matrix as JSON. Mutation-likelihood runs save the full per-position × 20-amino-acid logit matrix as JSON. Both are ready to load into a notebook, a clustering pipeline, or your own downstream scripts.
Raw vectors and raw likelihood matrices are never inlined into the chat — they live in the dataset, which keeps the chat fast and your results portable.
Pricing and limits
| Limit | Value |
|---|
| Concurrent runs per user | 3, shared with other GPU-class jobs |
| Minimum credit balance to start | 5,000 credits |
| Billing rate | 10 credits per second of actual compute, rounded up to the nearest minute |
| Dedup window | Identical input within 10 minutes returns the cached result at no extra cost |
Enabling models in your workspace
Protein embeddings and mutation likelihoods are managed per model in your capability settings, so you can turn on exactly the ones you need:
- AbLang-2 — Embeddings
- AbLang-2 — Mutation likelihoods
- ESM-2 — Embeddings
- ESM-2 — Mutation likelihoods
Enable the capabilities you need; leave off the ones you do not. Disabled capabilities are not offered to MIP as tools, which keeps your chat responses focused on the models you care about.
When to use which in a single workflow
A common pattern is combining the two model families across a multi-step analysis:
- Embed a batch of candidates to cluster them and pick representatives.
- Score mutation likelihoods on the representatives to find tolerant positions.
- Pick a small set of mutations at the top-scoring positions for wet-lab testing.
For an antibody workflow, replace ESM-2 with AbLang-2 in both steps. For generic proteins, ESM-2 is the right choice for both.
Mutation likelihoods are a ranking signal, not a guarantee. Use them to prioritize which positions to test, not to predict functional outcomes directly. Pair them with structure prediction to sanity-check that high-likelihood mutations sit in plausible structural contexts. Both ESM-2 and AbLang-2 are computational models. The outputs are ranking signals and feature representations, not experimental measurements. Treat them as hypothesis-generating inputs to your downstream workflow.

